ct_kmeans Centroid-based clustering
PRTools and ClusterTools should be in the path.
Goto ClusterTools examples for a review of all examples.
Download the m-file from here.
Contents
Prepare environment
prtime(10) % restrict iterative optimisation to 10s delfigs % delete existing figures randreset ; % takes care of reproducability prwarning(2) % show warnings
Define a dataset
we use some standard routines to create 8 two-dimensional clusters. They are labeld to compare cluster results with the desired labeling.A
m = 1000;
a = gendatclust2(m);
scattern(a); axis equal
title(getname(a));
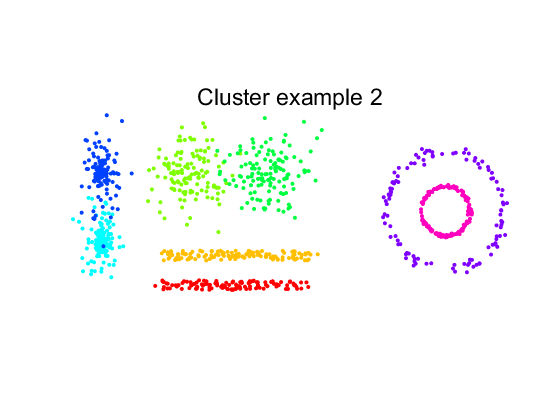

Run two clustering procedures after removing the label information.
x = +a; % remove labels K = [2 3 5 8 12 20 30 50 100]; % desired cluster sizes procs = {'KMeans','KCentres'}; err_I = cell(1,2); err_II= cell(1,2); figure; for i=1:2 subplot(2,2,i); lab = x*clustk(K,procs{i}); % execute KMeans or KCentres scatn(lab(:,4),x,procs{i}); % scatterplot markcols(1); % show distinguishable colors colors axis equal; err_I{i} = clusteval(lab,a,'actl'); err_II{i} = clusteval(lab,a,'roc'); end subplot(2,2,3); h1 = plote(err_I,10,'legend',procs); subplot(2,2,4); h2 = plote(err_II,10,'legend',procs);
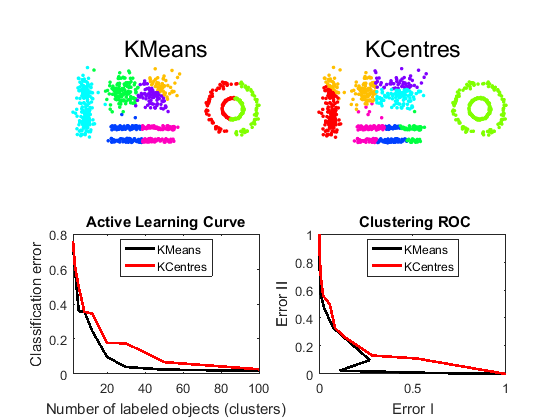
Comments
KMeans clustering looks somewhat better in the scatterplots. The bottom left curves show that in comparison with the desired labels (figure 1) this is supported by active labeling by which entire clusters are assigned to the label of their mediod or centre. (A cluster mean has no label, so the label of the nearest object, the cluster mediod, is used).
The bottom right figure shows the trade-off between the two types of errors as a function of the number of clusters. Error I is the fraction of object pairs that has been erroneously assigned to the same cluster. Error II is the fraction of object pairs that has been erroneously assigned to different clusters.